张长生 男 博导 南海海洋研究所
电子邮件:czhang@scsio.ac.cn;czhang2006@gmail.com
通信地址:广州市新港西路164号
邮政编码:510301
研究领域和科研项目
研究领域
围绕海洋微生物源药物分子的发现、生物合成、代谢工程和开发应用开展研究,探究多组学指导下海洋微生物源药物分子发现的新方法,解析复杂的生物合成途径和独特的酶催化机制,开展药物分子的生物活性和成药性评价研究,将合成生物学和代谢工程应用于海洋微生物源药物分子开发应用的实践和创新。
1. 生物大数据指导下的海洋微生物活性物质的发现。基于生物大数据,探究组学指导下的海洋微生物源药物分子发现的新方法,从中发现具有应用前景的先导分子,开展生物活性和成药性评价研究。
2. 药源分子生物合成及新功能酶的作用机制研究。阐明新药物分子的生物合成途径,揭示药物分子生物合成中包含的独特酶学机制和结构基础,定向改造开发具有高效催化功能的酶。
3. 合成生物学与代谢工程的实践和创新。采用合成生物学和代谢工程的技术手段,实现有应用前景的药物分子的产量提升和结构优化,为实现海洋微生物源药物分子的绿色制造奠定基础。
科研项目
国家重点研发计划项目,2023.12-2028.11
国家自然科学基金委
重大项目(参与)2022-2026
杰出青年基金 2012-2015
重点项目 2017-2021
重点国际(地区)合作研究项目 2019-2023
面上项目(3)
广州市南沙区重点领域科技计划项目 2023.12-2026.11
广东省海洋经济发展(海洋六大产业)专项资金项目 2021.4-2023.3
中国科学院王宽诚率先人才计划“卢嘉锡国际团队项目” 2021-2023
海南省科技厅重大项目 2020.12-2023.12
国家重点基础研究发展计划(973计划)课题 2010-2014
中科院前沿领域重点项目 2016-2022
重要方向性项目 2009-2011
中科院先导专项重点任务 2013-2017
广东省海洋经济创新发展区域示范项目,课题 2013-2017
广东省自然基金重大培育项目 2015-2018
广东省特支计划人才项目(2) 2015-2019
广东省渔港建设和渔业产业发展专项 2016-2018
代表性文章
Liu et al Journal of the American Chemical Society, 2023, 145 (50): 27886–27899. DOI: 10.1021/jacs.3c11772.
Jiang et al Angewandte Chemie International Edition, 2023, 62 (51): e202310728. DOI: 10.1002/anie.202310728.
Tan et al Angewandte Chemie International Edition, 2023, 62 (27): e202302043. DOI: 10.1002/anie.202302043.
Yang et al Nature Communications, 2022, 13: 5386. DOI: 10.1038/s41467-022-33131-0.
De et al Nature Communications, 2022, 13: 4896. DOI: 10.1038/s41467-022-32641-1.
Zhang et al Science China Chemistry, 2021, 64 (10): 1736–1742. DOI: 10.1007/s11426-021-1072-6.
Zhu et al Angewandte Chemie International Edition, 2020, 59 (33): 14065-14069. DOI: 10.1002/anie.202004978.
Huang et al Nature Communications, 2018, 9 (1): 2088. DOI: 10.1038/s41467-018-04487-z.
Zhang et al Science, 2006, 313(5791): 1291-1294. DOI: 10.1126/science.1130028.
招生信息
招生专业
070703-海洋生物学
070321-化学生物学
071005-微生物学
招生方向
海洋生物学、海洋天然产物化学、海洋微生物学
教育背景
1994-09--今 华东理工大学 硕士
1990-08--今 上海交通大学 学士
工作经历
2020/07 -- 今 中国科学院南海海洋研究所,副所长
2015/04 -- 2020/07 中国科学院南海海洋研究所,所长助理
2012/08 -- 2020/04 中国科学院南海海洋研究所,科研与规划处处长
2008/03 -- 今 中国科学院南海海洋研究所, 研究员
2003/01 -- 2008/03 美国威斯康星大学药学院博士后,assistant Scientist
1997/07 -- 1998/08 上海太平洋生物高科技有限公司,助理研究员
教授课程
获得荣誉
2024年: Editor, Marine Life Science & Technology
2023年: 2022年度中国海洋与湖沼十大科技进展
2022年: Associate Editor,Microbiome
2022年: Associate Editor,Environmental Microbiome
2021年: 中国科学院南海海洋研究所优秀研究生指导教师荣誉称号
2021年: 英国皇家化学学会会士(Fellow of Royal Society of Chemistry, FRSC)
2020年: 中国科学院南海海洋研究所优秀研究生指导教师荣誉称号
2020年: 编委,Molecules
2019年: Editorial board member, Natural Product Reports
2018年: 2018年度"中国科学院优秀导师奖"
2018年: 2018年度中国科学院广州教育基地"优秀研究生导师"
2018年: 中国科学院南海海洋研究所优秀研究生指导教师荣誉称号
2017年: 享受“国务院政府特殊津贴”
2016年: 国家科技创新领军人才
2016年: 第四届曾呈奎海洋科技奖--青年科技奖,中国海洋湖沼学会
2015年: 广东特支计划“杰出人才(南粤百杰)”
2014年: 有突出贡献中青年专家
2014年: 中青年科技创新领军人才
2014年: 广东特支计划”领军人才”
2013年: 择优终期评估”优秀”,中国科学院
2012年: 全国优秀科技工作者,中国科协
2011年: 国家杰出青年基金获得者(终期评估优秀),国家自然科学基金委员会
2009年: 择优支持入选者,中国科学院
2005年: “First Midwest Carbohydrate Symposium”, Anatrace Best Poster Award
出版信息
# (co-first author), * (corresponding author)
2024
180. Liangliang Bai,# Linping Wu,# Changsheng Zhang, Zhiwen Liu, Liang Ma, Jing Ni, Dezhen He, Mingxuan Zhu, Shaoyong Peng, Xiaoxia Liu, Huichuan Yu, Yuhe Lei, Yanxin Luo, Yu Zhang, Xiaolin Wang,* Gang Wei,* Yingjie Li.* Replenishment of mitochondrial Na+ and H+ by ionophores potentiates cutaneous wound healing in diabetes. Materials Today Bio, 2024, DOI: 10.1016/j.mtbio.2024.101056. Available online 16 April 2024;
179. Yanqin Li,# Mengjing Cong,# Xiufeng Zhang, Yiguang Zhu, Wenjun Zhang, Yonghong Liu, Changsheng Zhang,* Junfeng Wang,* and Yan Yan.* An Enzymatic Carbon-Carbon Bond Cleavage and Aldol Reaction Cascade Converts an Angular Scaffold into the Linear Tetracyclic Core of Ochraceopones. Angewandte Chemie International Edition, 2024, 63, e202403365. DOI: 10.1002/anie.202403365. Accepted manuscript online: 07 March 2024; Version of Record online: 26 March 2024;
178. 姚景龙,陈海涛,张长生*,杨敏,王卫强,魏源送,潘刚,王亚炜,罗耀,钟慧,张镇秋。践行“一带一路”新发展理念,推动“中国-斯里兰卡联合科教中心”高质量发展。中国科学院院刊,2024, 39 (2): 381-386. DOI: 10.16418/j.issn.1000-3045.20230908002. Published 21 Feb 2024.
177. Youzhe Chen,# Hongbo Jin,# Weiliang Xiong, Zhuangjie Fang, Weiguang Sun,Yiguang Zhu, Liping Zhang, Yonghui Zhang, Wenjun Zhang,* Changsheng Zhang.* Discovery of Aburatubolactams Reveals Biosynthetic Logic for Distinct 5/5-Type Polycyclic Tetramate Macrolactams. Organic Letters, 2024, 26 (8): 1677–1682. DOI: 10.1021/acs.orglett.4c00168. Publication Date: February 16, 2024; Published in issue 1 March 2024.

176. Wei Liu,# Shilan Zhai,# Liang Ma,* Liping Zhang, Yuchan Chen, Zhiyong Liu, Tianyu Zhang, Weimin Zhang, Changsheng Zhang,* Wenjun Zhang.* Expanding the Chemical Diversity of Grisechelins via Heterologous Expression. Journal of Natural Products, 2024, 87 (2): 371-380. DOI: 10.1021/acs.jnatprod.3c01132. Published online 1 February 2024; Published in issue 23 February 2024.

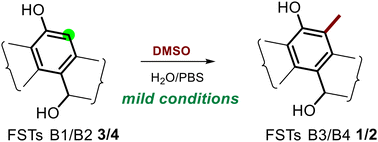
175. Bidhan Chandra De, Chunfang Yang,
Chunshuai Huang, Changsheng Zhang, and Wenjun Zhang.* Non-enzymatic synthesis of C-methylated fluostatins: discovery and reaction mechanism. Organic & Biomolecular Chemistry, 2024, 22 (6): 1152-1156. DOI: 10.1039/d3ob01920a. first published on 03 Jan 2024; advance article 12 Jan 2024; in issue 7 Feb 2024.
174. Bin Tan, Qingbo Zhang, Liping Zhang, Yiguang Zhu, Changsheng Zhang.* Naturally Occurring and Widespread Resistance to Bioactive Natural Products. ChemMedChem, 2024, 19 (3): e202300619. DOI: 10.1002/cmdc.202300619 (invited review). Accepted manuscript online: 16 December 2023; Version of Record online: 12 January 2024; Issue Online: 01 February 2024.
2023
173. Kai Liu,# Jinyan Zhang,# Guangtao Zhang,#* Liping Zhang, Zhen Meng, Liang Ma, Wenjun Zhang, Weiliang Xiong, Yiguang Zhu, Binju Wang,* Changsheng Zhang.* Deciphering Deoxynybomycin Biosynthesis Reveals Fe(II)/α-Ketoglutarate-Dependent Dioxygenase-Catalyzed Oxazoline Ring Formation and Decomposition. Journal of the American Chemical Society, 2023, 145 (50): 27886–27899. DOI: 10.1021/jacs.3c11772. Publication online 6 December, 2023. (